Microenvironment signals are potent determinants of cell fate and arbiters of tissue homeostasis, however understanding how different microenvironment factors coordinately regulate cellular phenotype has been experimentally challenging. Here we used a high-throughput microenvironment microarray comprised of 2640 unique pairwise signals to identify factors that support proliferation and maintenance of primary human mammary luminal epithelial cells. Multiple microenvironment factors that modulated luminal cell number were identified, including: HGF, NRG1, BMP2, CXCL1, TGFB1, FGF2, PDGFB, RANKL, WNT3A, SPP1, HA, VTN, and OMD. All of these factors were previously shown to modulate luminal cell numbers in painstaking mouse genetics experiments, or were shown to have a role in breast cancer, demonstrating the relevance and power of our high-dimensional approach to dissect key microenvironmental signals. RNA-sequencing of primary epithelial and stromal cell lineages identified the cell types that express these signals and the cognate receptors in vivo. Cell-based functional studies confirmed which effects from microenvironment factors were reproducible and robust to individual variation. Hepatocyte growth factor (HGF) was the factor most robust to individual variation and drove expansion of luminal cells via cKit+ progenitor cells, which expressed abundant MET receptor. Luminal cells from women who are genetically high risk for breast cancer had significantly more MET receptor and may explain the characteristic expansion of the luminal lineage in those women. In ensemble, our approach provides proof of principle that microenvironment signals that control specific cellular states can be dissected with high-dimensional cell-based approaches
Main navigation
- Main
-
About AAUP
Image


Give university
For those who wish to donate to the university, you can make your donations by filling out the donation form
-
Admissions & Academics
- Deanship of Admission and Registration
- Programs
- Prospective Student Admission
- Faculties
- Academic Quality
- Bridging to Bachelor's Degree
- Transfer to AAUP
- Visiting student
- Scholarships and Financial Aid
- Mobility and External Scholarships
- Tuition Fees and Costs
- E-Learning
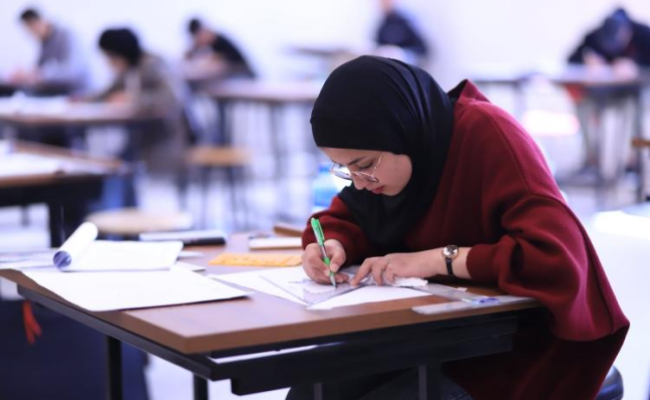
- English Language Placement Test
- Academic Schedule
Image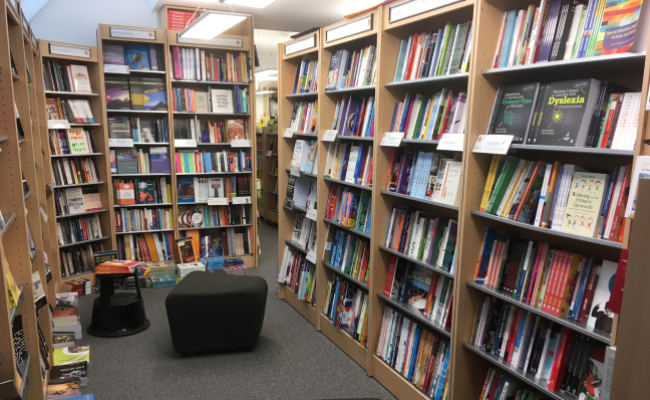

Arab American University Library
A distinguished library at the local and international levels by providing all electronic and paper information resources.
-
University life
Image
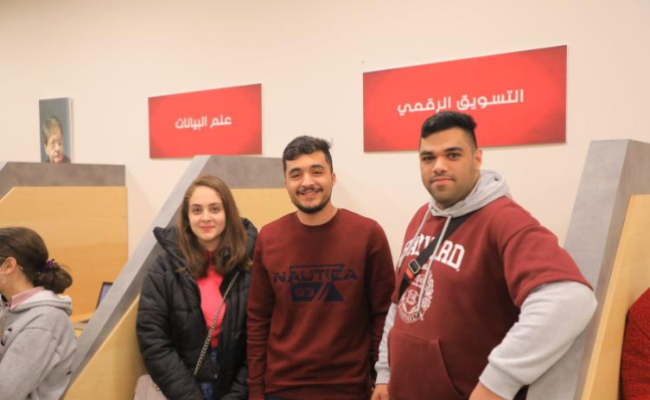


Inspiring success stories from the Arab American University
Success represents the pinnacle of persistence in the face of failure. So, these are the success stories we will share with you that reveal the path taken by some of our university’s graduates.
-
University centers
- Center for Research and Polling
- Climate Change Center
- Conflict Studies Research Center
- Continuing Education Center
- Dental Center
- E-Learning Center
- Hassib Sabbagh Center of Excellence
- Heart Center
- Language Center
- Medical Center - Ramallah
- National Digital Transformation and Artificial Intelligence Center
- Prosthetics and Orthotics Factory
- Simulation Center
Image
Medical Rehabilitation Complex
-
Research
Image


Arab American University Awards
As part of the university’s efforts to support and encourage scientific research, there are a set of awards to encourage researchers to excel in their original and valuable scientific production
-
News & Media
Image


AAUP ACADEMIC CALENDAR
An agenda to show the the important dates such as semester dates, registration, exams, activities, events, and holidays.
